In connection with our projects, the Sabeti Lab has created a number of open source software tools:
Our Software
Composite of Multiple Signals (“CMS”): tests for selection in meiotically recombinant populations
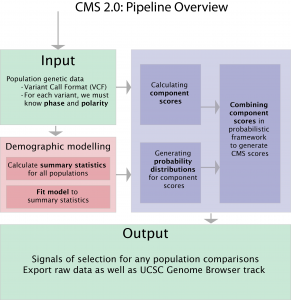
Documentation
Source code
Cosi2: a fast coalescent simulator with selection
Installation instructions
Documentation
Source code
viral-ngs: genomic analysis scripts and pipelines for viral sequencing
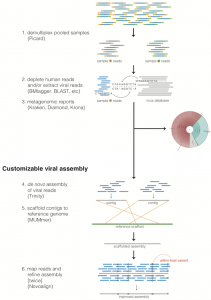
Documentation
Source code
Mirador: a tool to identify new hypothesis in complex datasets through visual exploration
Home page
Documentation
Source code
Mirador Open Data Competition
Information
National Health and Nutrition Examination Survey Data and Scripts
GitHub repository
Ebola CARE prognosis prediction and treatment recommendation app

Home page
Google Play page
Source code
sabetilab-remote-config: configuration files for systems deployed at our African field sites
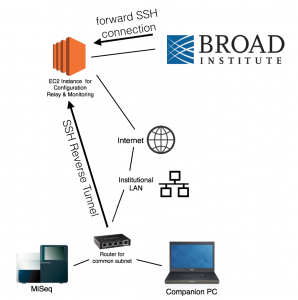
Source code
Contributions
Members of the Sabeti Lab are also contributors to a number of external open source projects with relevance to the biological sciences:
bioconda: a channel for the conda package manager specializing in bioinformatics software

Documentation
Snakemake: a scalable bioinformatics workflow engine (in Python)
Documentation
Processing: a flexible software sketchbook and a language for learning how to code within the context of the visual arts.

